Transforming raw data into ICER plots using the easyicer package
easyicer-vignette.RmdIntroduction
This vignette is a tutorial on how to seamlessly create incremental cost-effectiveness ratio (ICER) analyses, including plots, by making use of all the functions in this package. This package was designed in a pipeline structure so that the end-user could cohesively deploy the package functions in a step-wise manner to arrive at a final cost-effectiveness analysis (CEA) with ICER plots. For instructive purposes, this vignette will be organized into 3 use-cases that can be integrated as a single workflow or deployed standalone. The target domain for this package is health technology assessment and economics, where CEAs are a common tool for evaluating the risk-benefit of new health interventions for implementation.
Data
The data used in this tutorial is a modified dataset obtained from the authors’ research project. The full dataset is not publicly available. The modified dataset used herein can be located in the GitHub repository for this package. For the purpose of this tutorial, we will assign the dataset to an object.
# Load the "here" package to access the .csv file in the data folder if you are in the source package repository.
easyicer_data <- read.csv(here::here("data/easyicer_data.csv"))
dim(easyicer_data)
#> [1] 828 8
head(easyicer_data, n = 10)
#> cohort scr_age ppt_rate scr_mod year Cost_of_screen_x_reen_positives
#> 1 1973 50 0.18 FIT 2015 27458883
#> 2 1973 50 0.18 FIT 2016 27805623
#> 3 1973 50 0.18 FIT 2017 27650384
#> 4 1973 50 0.18 FIT 2018 16568149
#> 5 1973 50 0.18 FIT 2019 16598985
#> 6 1973 50 0.18 FIT 2020 17469883
#> 7 1973 50 0.18 FIT 2021 17340106
#> 8 1973 50 0.18 FIT 2022 17281809
#> 9 1973 50 0.18 FIT 2023 22268950
#> 10 1973 50 0.18 FIT 2024 29240148
#> Cost_of_clinic_x_ow_up_protocol Health_adjusted_person_years
#> 1 1660290 2132296
#> 2 3498682 2141253
#> 3 3764434 2149907
#> 4 6989066 2151707
#> 5 7421521 2149004
#> 6 7601313 2147614
#> 7 8097227 2145495
#> 8 8479750 2142464
#> 9 7999188 2138152
#> 10 8926254 2132390The dataset contains 8 variables with 828 rows of observations. This data was obtained from a microsimulation model projecting lifetime health and economic outcomes of average-risk colorectal cancer screening simulations in pre-defined Canadian birth cohorts. There are 4 predictor variables, or characteristics (cohort, scr_age, ppt_rate, scr_mod, year), and 3 outcome variables (including both cost outcomes and a health outcome). See below for full variable definitions.
Variable Dictionary
cohort: A simulated birth cohort in the Canadian population with modeled adenoma risk relative to the age group. Example: 1973 represents a cohort of individuals born between 1973 and 1977.
scr_age: The modeled age of screening initiation for each scenario. Example: 50 means that simulated individuals began screening at age 50.
ppt_rate: The modeled participation rate for colorectal cancer screening in the screening scenario. Example: 0.18 = 18% of the cohort underwent screening.
scr_mod: The modality of the screening test. Either a fecal immunochemical test (FIT) or a colonoscopy (f10q).
year: The corresponding year that outcomes were simulated for.
Cost_of_screen_x_reen_positives: The total cost of screening, including follow-up screening of positive cases.
Cost_of_clinic_x_ow_up_protocol: The cumulative cost incurred by year from clinical diagnoses and follow-up procedures.
Health_adjusted_person_years: The estimated cumulative life years of all individuals based on individually ascribed health utility scores. Herein defined as quality-adjusted life years (QALY).
Use-Case 1: Calculating cumulative outputs for cost and QALY
outcomes with summed_output()
Often times with health economic data, we might get a large volume of
raw and unprocessed outputs. This is particularly the case for
simulation models that generate and display highly granular information.
Often times, the individual outputs are meaningless unless they are
transformed in some way (e.g., calculating a cumulative value) This is
where it is helpful to deploy the summed_output() function,
especially when this process is iterative. The
summed_output() function calculates the sum of a numeric
column in the dataset to arrive at a cumulative value. With this
dataset, this is valuable because we need to calculate the sum of each
output variable (costs and QALY) since they were originally calculated
per year. This function also allows the end-user to group by and filter
specific characteristics. For example, in our data, we have multiple
birth cohorts, screening ages and screening modalities that we want to
group by before calculating cumulative values. Lastly, the function can
handle any NA values that might otherwise disrupt the function
calculation.
# Set a filter parameter to call in filter_vars
ppt18 <- easyicer_data$ppt_rate == 0.18
# Calculate the cumulative values for the cost outcome
cumulative_cost <- summed_output(
data = easyicer_data,
group_vars = c("scr_mod", "scr_age"),
sum_var = "Cost_of_screen_x_reen_positives",
filter_vars = ppt18
)
# Calculate the cumulative values for the health outcome
cumulative_qaly <- summed_output(
data = easyicer_data,
group_vars = c("scr_mod", "scr_age"),
sum_var = "Health_adjusted_person_years",
filter_vars = ppt18
)
print(cumulative_cost)
#> # A tibble: 4 × 3
#> # Groups: scr_mod [2]
#> scr_mod scr_age cumulative
#> <chr> <int> <dbl>
#> 1 FIT 45 2694728140.
#> 2 FIT 50 2517849263.
#> 3 f10q 45 3420224126.
#> 4 f10q 50 3312806274.
print(cumulative_qaly)
#> # A tibble: 4 × 3
#> # Groups: scr_mod [2]
#> scr_mod scr_age cumulative
#> <chr> <int> <dbl>
#> 1 FIT 45 205541825.
#> 2 FIT 50 205524017.
#> 3 f10q 45 205610208.
#> 4 f10q 50 205601137.Now we have arrived at our necessary outputs to calculate ICERs. Did
you notice that this function is a general sum function with the ability
to sum by groups or filtered datasets? This function is not strictly
intended for calculating ICERs and is versatile with other purposes, but
it makes iterative sum calculations very convenient - especially when
calculating ICERs. Note that we did not specifically call
na.rm = TRUE, this is because it is set to
TRUE as default so if it is not called in the function it
will automatically remove NA values.
Use-Case 2: Calculating ICERs from cumulative cost and QALY inputs
with icercalc()
Before we can calculate ICERs, we need to transform the data
structure so that the values are recognized in the correct order by the
icercalc() function. This is because with ICERs, there are
reference values that correspond with a reference or comparator
scenario/intervention. In the context of our dataset, this refers to
scr_age = 50 and scr_mod = FIT (FIT 50), which
is the current colorectal cancer screening strategy in Canada. In order
to properly calculate the ICERs, the function expects a matrix structure
as illustrated:
ref new
cost n00 n01
qaly n10 n11Once you have created a matrix, you might realize that the order of
values in the matrix is not correct. Instead of manually re-coding the
matrix, you can specify rev = as "neither",
"rows", "columns" or "both" to
reverse the order of the columns, rows, or columns and rows. By default
rev = "neither" so you do not need to specify the
rev argument if the matrix is correct. For instructive
purposes, the rev argument will be showcased below.
# Create matrix of cost and QALY values for each scenario
icer_matrix_fit45 <- matrix(c(
cumulative_cost$cumulative[2],
cumulative_cost$cumulative[1],
cumulative_qaly$cumulative[2],
cumulative_qaly$cumulative[1]),
nrow = 2,
ncol = 2,
byrow = TRUE)
# Calculate ICERs using matrix
icer_fit45 <- icercalc(data = icer_matrix_fit45)
#> [1] "Modified matrix:"
#> [,1] [,2]
#> [1,] 2517849263 2694728140
#> [2,] 205524017 205541825
#>
#> Incremental Cost:
#> 176878878
#> Incremental QALY:
#> 17807.78
#> Incremental cost-effectiveness ratio (ICER):
#> 9932.676
# Re-iteration for COL 45 scenario
icer_matrix_col45 <- matrix(c(
cumulative_cost$cumulative[2],
cumulative_cost$cumulative[3],
cumulative_qaly$cumulative[2],
cumulative_qaly$cumulative[3]),
nrow = 2,
ncol = 2,
byrow = TRUE)
icer_col45 <- icercalc(data = icer_matrix_col45)
#> [1] "Modified matrix:"
#> [,1] [,2]
#> [1,] 2517849263 3420224126
#> [2,] 205524017 205610208
#>
#> Incremental Cost:
#> 902374863
#> Incremental QALY:
#> 86190.82
#> Incremental cost-effectiveness ratio (ICER):
#> 10469.5
# # Re-iteration for COL 50 scenario
icer_matrix_col50 <- matrix(c(
cumulative_cost$cumulative[4], # The columns of the matrix are purposefully in inverse order to demonstrate rev argument.
cumulative_cost$cumulative[2],
cumulative_qaly$cumulative[4],
cumulative_qaly$cumulative[2]),
nrow = 2,
ncol = 2,
byrow = TRUE)
icer_col50 <- icercalc(data = icer_matrix_col50, rev = "columns")
#> [1] "Modified matrix:"
#> [,1] [,2]
#> [1,] 2517849263 3312806274
#> [2,] 205524017 205601137
#>
#> Incremental Cost:
#> 794957011
#> Incremental QALY:
#> 77119.96
#> Incremental cost-effectiveness ratio (ICER):
#> 10308.06The output of icercalc() displays the final matrix used
in the ICER calculation, the incremental cost output (difference in cost
between the new and reference scenarios), the incremental QALY
(difference in QALY between new and reference scenarios) and the ICER.
The component cost and QALY outcomes are displayed because if the end
user works with the entire easyicer pipeline, both values will be
required for the icerplot() function.
Use-Case 3: Creating ICER plots from calculated ICERs with
icerplot()
The last stage of a basic CEA is to plot the ICERs along a coordinate
system. This is helpful for visually comparing how cost-effective
multiple interventions are relative to the reference intervention. The
icerplot() function creates a scatterplot based on a
dataframe with at least two numeric columns (representing cost and QALY
outcomes). An additional and optional names argument is
implemented which creates legend labels if the ICER values have
corresponding names.
# Create a dataframe (if needed) with cost and QALY derivatives of ICER values.
icer_coordinates <- data.frame(
intervention = c("FIT 45", "COL 45", "COL 50"),
QALY = c(17807.78, 86190.82, 77119.96),
Cost = c(176878878, 902374863, 794957011))
icerplot(data = icer_coordinates, x = "QALY", y = "Cost", names = "intervention")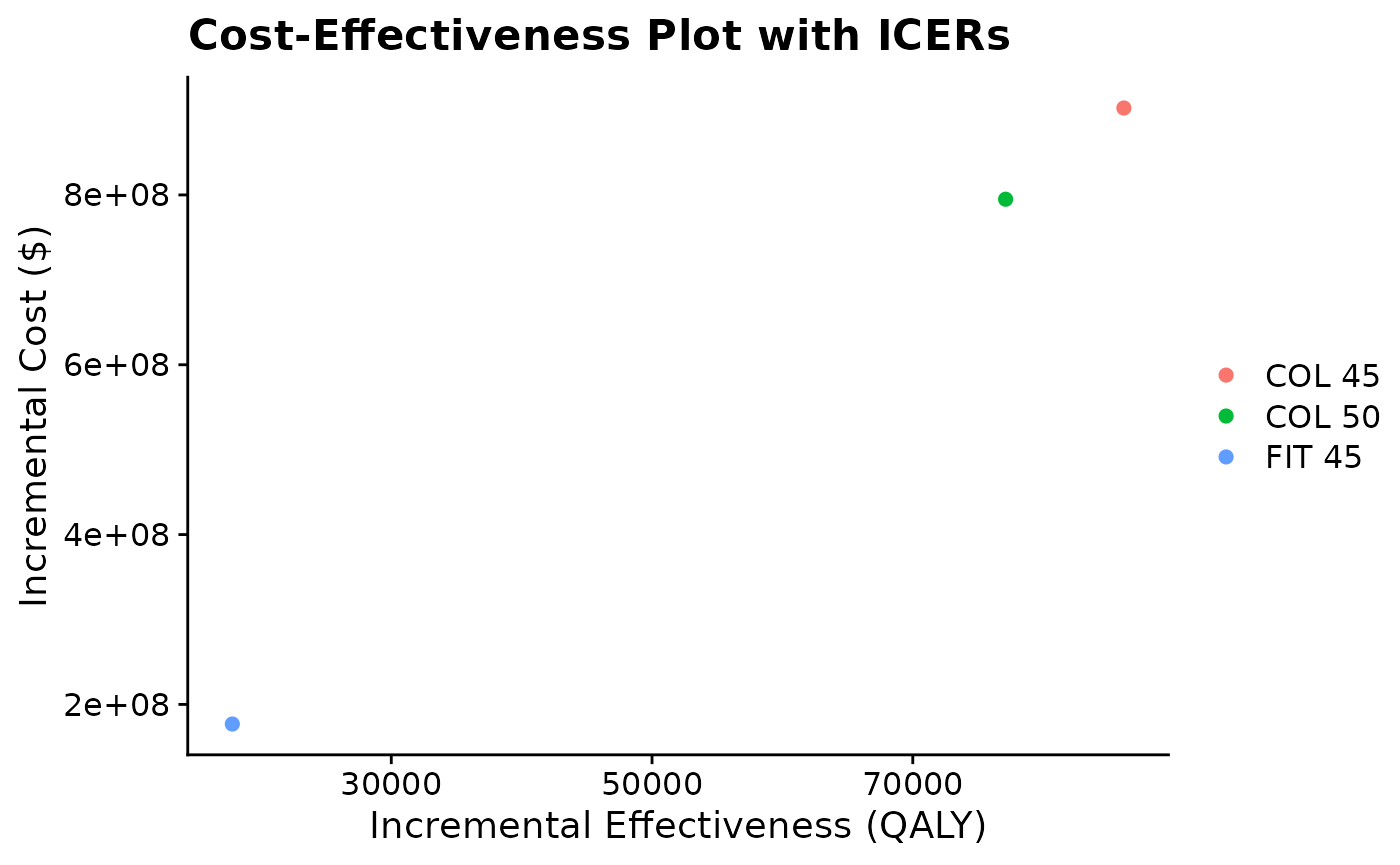
Notice that you do not need the ICER outputs from
icercalc() to use this function. In fact, the ICER values
calculated with icercalc() will not work since both the
component costs and QALY outcomes are required in the dataframe, which
is why icercalc() provides both incremental costs and QALYs
as well as the ICER. This highlights that these functions can be used in
an integrated manner or independently based on the end user’s needs and
data structures.